Swap in one other LLM
There are a number of methods to run the identical activity with a special mannequin. First, create a brand new chat object with that completely different mannequin. Right here’s the code for trying out Google Gemini 3 Flash Preview:
my_chat_gemini <- chat_google_gemini(mannequin = "gemini-3-flash-preview")Then you possibly can run the duty in one in every of 3 ways.
1. Clone an current activity and add the chat as its solver with $set_solver():
my_task_gemini <- my_task$clone()
my_task_gemini$set_solver(generate(my_chat_gemini))
my_task_gemini$eval(epochs = 3)2. Clone an current activity and add the brand new chat as a solver if you run it:
my_task_gemini <- my_task$clone()
my_task_gemini$eval(epochs = 3, solver_chat = my_chat_gemini)3. Create a brand new activity from scratch, which lets you embrace a brand new title:
my_task_gemini <- Activity$new(
dataset = my_dataset,
solver = generate(my_chat_gemini),
scorer = model_graded_qa(
partial_credit = FALSE,
scorer_chat = ellmer::chat_anthropic(mannequin = "claude-opus-4-6")
),
title = "Gemini flash 3 preview"
)
my_task_gemini$eval(epochs = 3)Be sure you’ve set your API key for every supplier you need to take a look at, until you’re utilizing a platform that doesn’t want them, resembling native LLMs with ollama.
View a number of activity runs
When you’ve run a number of duties with completely different fashions, you need to use the vitals_bind() perform to mix the outcomes:
both_tasks <- vitals_bind(
gpt5_nano = my_task,
gemini_3_flash = my_task_gemini
)Instance of mixed activity outcomes operating every LLM with three epochs.
Sharon Machlis
This returns an R knowledge body with columns for activity, id, epoch, rating, and metadata. The metadata column accommodates a knowledge body in every row with columns for enter, goal, consequence, solver_chat, scorer_chat, scorer_metadata, and scorer.
To flatten the enter, goal, and consequence columns and make them simpler to scan and analyze, I un-nested the metadata column with:
library(tidyr)
both_tasks_wide <- both_tasks |>
unnest_longer(metadata) |>
unnest_wider(metadata)I used to be then in a position to run a fast script to cycle via every bar-chart consequence code and see what it produced:
library(dplyr)
# Some outcomes are surrounded by markdown and that markdown code must be eliminated or the R code will not run
extract_code <- perform(textual content) ```", "", textual content)
textual content <- trimws(textual content)
textual content
# Filter for barchart outcomes solely
barchart_results <- both_tasks_wide |>
filter(id == "barchart")
# Loop via every consequence
for (i in seq_len(nrow(barchart_results))) {
code_to_run <- extract_code(barchart_results$consequence[i])
rating <- as.character(barchart_results$rating[i])
task_name <- barchart_results$activity[i]
epoch <- barchart_results$epoch[i]
# Show information
cat("n", strrep("=", 60), "n")
cat("Activity:", task_name, "| Epoch:", epoch, "| Rating:", rating, "n")
cat(strrep("=", 60), "nn")
# Attempt to run the code and print the plot
tryCatch(
{
plot_obj <- eval(parse(textual content = code_to_run))
print(plot_obj)
Sys.sleep(3)
},
error = perform(e) {
cat("Error operating code:", e$message, "n")
Sys.sleep(3)
}
)
}
cat("nFinished displaying all", nrow(barchart_results), "bar charts.n")Take a look at native LLMs
That is one in every of my favourite use instances for vitals. At the moment, fashions that match into my PC’s 12GB of GPU RAM are reasonably restricted. However I’m hopeful that small fashions will quickly be helpful for extra duties I’d love to do regionally with delicate knowledge. Vitals makes it simple for me to check new LLMs on a few of my particular use instances.
vitals (through ellmer) helps ollama, a preferred manner of operating LLMs regionally. To make use of ollama, obtain, set up, and run the ollama utility, and both use the desktop app or a terminal window to run it. The syntax is ollama pull to obtain an LLM, or ollama run to each obtain and begin a chat should you’d like to verify the mannequin works in your system. For instance: ollama pull ministral-3:14b.
The rollama R package deal enables you to obtain an area LLM for ollama inside R, so long as ollama is operating. The syntax is rollama::pull_model("model-name"). For instance, rollama::pull_model("ministral-3:14b"). You’ll be able to take a look at whether or not R can see ollama operating in your system with rollama::ping_ollama().
I additionally pulled Google’s gemma3-12b and Microsoft’s phi4, then created duties for every of them with the identical dataset I used earlier than. Be aware that as of this writing, you want the dev model of vitals to deal with LLM names that embrace colons (the subsequent CRAN model after 0.2.0 ought to deal with that, although):
# Create chat objects
ministral_chat <- chat_ollama(
mannequin = "ministral-3:14b"
)
gemma_chat <- chat_ollama(
mannequin = "gemma3:12b"
)
phi_chat <- chat_ollama(
mannequin = "phi4"
)
# Create one activity with ministral, with out naming it
ollama_task <- Activity$new(
dataset = my_dataset,
solver = generate(ministral_chat),
scorer = model_graded_qa(
scorer_chat = ellmer::chat_anthropic(mannequin = "claude-opus-4-6")
)
)
# Run that activity object's evals
ollama_task$eval(epochs = 5)
# Clone that activity and run it with completely different LLM chat objects
gemma_task <- ollama_task$clone()
gemma_task$eval(epochs = 5, solver_chat = gemma_chat)
phi_task <- ollama_task$clone()
phi_task$eval(epochs = 5, solver_chat = phi_chat)
# Flip all these outcomes right into a mixed knowledge body
ollama_tasks <- vitals_bind(
ministral = ollama_task,
gemma = gemma_task,
phi = phi_task
)All three native LLMs nailed the sentiment evaluation, and all did poorly on the bar chart. Some code produced bar charts however not with axes flipped and sorted in descending order; different code didn’t work in any respect.
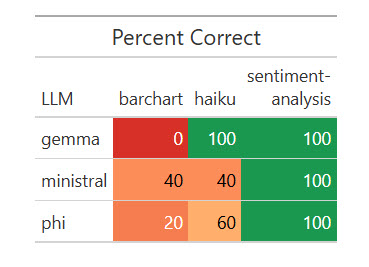

Outcomes of 1 run of my dataset with 5 native LLMs.
Sharon Machlis
R code for the outcomes desk above:
library(dplyr)
library(gt)
library(scales)
# Put together the information
plot_data <- ollama_tasks |>
rename(LLM = activity, activity = id) |>
group_by(LLM, activity) |>
summarize(
pct_correct = imply(rating == "C") * 100,
.teams = "drop"
)
color_fn <- col_numeric(
palette = c("#d73027", "#fc8d59", "#fc8d59", "#fee08b", "#1a9850"),
area = c(0, 20, 40, 60, 100)
)
plot_data |>
tidyr::pivot_wider(names_from = activity, values_from = pct_correct) |>
gt() |>
tab_header(title = "% Right") |>
cols_label(`sentiment-analysis` = html("sentiment-
evaluation")) |>
data_color(
columns = -LLM,
fn = color_fn
)It price me 39 cents for Opus to guage these native LLM runs—not a nasty cut price.
Extract structured knowledge from textual content
Vitals has a particular perform for extracting structured knowledge from plain textual content: generate_structured(). It requires each a chat object and an outlined knowledge sort you need the LLM to return. As of this writing, you want the event model of vitals to make use of the generate_structured() perform.
First, right here’s my new dataset to extract matter, speaker title and affiliation, date, and begin time from a plain-text description. The extra advanced model asks the LLM to transform the time zone to Jap Time from Central European Time:
extract_dataset <- knowledge.body(
id = c("entity-extract-basic", "entity-extract-more-complex"),
enter = c(
"Extract the workshop matter, speaker title, speaker affiliation, date in 'yyyy-mm-dd' format, and begin time in 'hh:mm' format from the textual content under. Assume the date 12 months makes essentially the most sense on condition that right now's date is February 7, 2026. Return ONLY these entities within the format {matter}, {speaker title}, {date}, {start_time}. R Bundle Growth in PositronrnThursday, January fifteenth, 18:00 - 20:00 CET (Rome, Berlin, Paris timezone) rnStephen D. Turner is an affiliate professor of knowledge science on the College of Virginia College of Knowledge Science. Previous to re-joining UVA he was a knowledge scientist in nationwide safety and protection consulting, and later at a biotech firm (Colossal, the de-extinction firm) the place he constructed and deployed scores of R packages. ",
"Extract the workshop matter, speaker title, speaker affiliation, date in 'yyyy-mm-dd' format, and begin time in Jap Time zone in 'hh:mm ET' format from the textual content under. (TZ is the time zone). Assume the date 12 months makes essentially the most sense on condition that right now's date is February 7, 2026. Return ONLY these entities within the format {matter}, {speaker title}, {date}, {start_time}. Convert the given time to Jap Time if required. R Bundle Growth in PositronrnThursday, January fifteenth, 18:00 - 20:00 CET (Rome, Berlin, Paris timezone) rnStephen D. Turner is an affiliate professor of knowledge science on the College of Virginia College of Knowledge Science. Previous to re-joining UVA he was a knowledge scientist in nationwide safety and protection consulting, and later at a biotech firm (Colossal, the de-extinction firm) the place he constructed and deployed scores of R packages. "
),
goal = c(
"R Bundle Growth in Positron, Stephen D. Turner, College of Virginia (or College of Virginia College of Knowledge Science), 2026-01-15, 18:00. OR R Bundle Growth in Positron, Stephen D. Turner, College of Virginia (or College of Virginia College of Knowledge Science), 2026-01-15, 18:00 CET.",
"R Bundle Growth in Positron, Stephen D. Turner, College of Virginia (or College of Virginia College of Knowledge Science), 2026-01-15, 12:00 ET."
)
)Beneath is an instance of outline a knowledge construction utilizing ellmer’s type_object() perform. Every of the arguments provides the title of a knowledge discipline and its sort (string, integer, and so forth). I’m specifying I need to extract a workshop_topic, speaker_name, current_speaker_affiliation, date (as a string), and start_time (additionally as a string):
my_object <- type_object(
workshop_topic = type_string(),
speaker_name = type_string(),
current_speaker_affiliation = type_string(),
date = type_string(
"Date in yyyy-mm-dd format"
),
start_time = type_string(
"Begin time in hh:mm format, with timezone abbreviation if relevant"
)
)Subsequent, I’ll use the chat objects I created earlier in a brand new structured knowledge activity, utilizing Sonnet because the decide since grading is simple:
my_task_structured <- Activity$new(
dataset = extract_dataset,
solver = generate_structured(
solver_chat = my_chat,
sort = my_object
),
scorer = model_graded_qa(
partial_credit = FALSE,
scorer_chat = ellmer::chat_anthropic(mannequin = "claude-sonnet-4-6")
)
)
gemini_task_structured <- my_task_structured$clone()
# It's essential add the kind to generate_structured(), that is not included when a structured activity is cloned
gemini_task_structured$set_solver(
generate_structured(solver_chat = my_chat_gemini, sort = my_object)
)
ministral_task_structured <- my_task_structured$clone()
ministral_task_structured$set_solver(
generate_structured(solver_chat = ministral_chat, sort = my_object)
)
phi_task_structured <- my_task_structured$clone()
phi_task_structured$set_solver(
generate_structured(solver_chat = phi_chat, sort = my_object)
)
gemma_task_structured <- my_task_structured$clone()
gemma_task_structured$set_solver(
generate_structured(
solver_chat = gemma_chat,
sort = my_object
)
)
# Run the evaluations!
my_task_structured$eval(epochs = 3)
gemini_task_structured$eval(epochs = 3)
ministral_task_structured$eval(epochs = 3)
gemma_task_structured$eval(epochs = 3)
phi_task_structured$eval(epochs = 3)
# Save outcomes to knowledge body
structured_tasks <- vitals_bind(
gemini = gemini_task_structured,
gpt_5_nano = my_task_structured,
ministral = ministral_task_structured,
gemma = gemma_task_structured,
phi = phi_task_structured
)
saveRDS(structured_tasks, "structured_tasks.Rds")It price me 16 cents for Sonnet to guage 15 analysis runs of two queries and outcomes every.
Listed here are the outcomes:
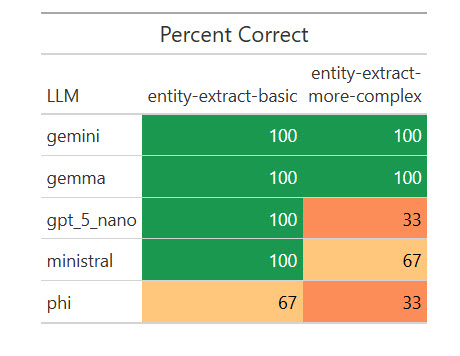
How numerous LLMs fared on extracting structured knowledge from textual content.
Sharon Machlis
I used to be stunned {that a} native mannequin, Gemma, scored 100%. I wished to see if that was a fluke, so I ran the eval one other 17 occasions for a complete of 20. Weirdly, it missed on two of the 20 primary extractions by giving the title as “R Bundle Growth” as an alternative of “R Bundle Growth in Positron,” however scored 100% on the extra advanced ones. I requested Claude Opus about that, and it stated my “simpler” activity was extra ambiguous for a much less succesful mannequin to know. Vital takeaway: Be as particular as doable in your directions!
Nonetheless, Gemma’s outcomes have been adequate on this activity for me to contemplate testing it on some real-world entity extraction duties. And I wouldn’t have identified that with out operating automated evaluations on a number of native LLMs.
Conclusion
If you happen to’re used to writing code that offers predictable, repeatable responses, a script that generates completely different solutions every time it runs can really feel unsettling. Whereas there are not any ensures on the subject of predicting an LLM’s subsequent response, evals can enhance your confidence in your code by letting you run structured assessments with measurable responses, as an alternative of testing through guide, ad-hoc queries. And, because the mannequin panorama retains evolving, you possibly can keep present by testing how newer LLMs carry out—not on generic benchmarks, however on the duties that matter most to you.

